Changes in the climate affect the survival and long-term viability of natural forests, therefore genomic information is key to understand the capacity of natural populations of tree species to adapt to new environmental conditions and diseases, and to develop effective conservation and management strategies. Research in our lab focuses on understanding the genomic and evolutionary basis of phenotypic trait variation, plant stress response, and adaptation to changing environments, using tools in genomics, transcriptomics, epigenomics and physiology.
Genomics of local adaptation to climate
Understanding the genomic basis of local adaptation is crucial to determine the potential of long lived woody species to withstand changes in their natural environments. Whole-genome studies were not feasible until very recently due to the absence of reference gymnosperm genomes, therefore previous studies were based on a small number of genes. Previous studies in our research group focused on dissecting the genomic architectures of local adaptation in several conifer species such as loblolly pine (De La Torre et 2019 GBE), Douglas fir ( De La Torre et al 2021 Genes), giant sequoia, coast redwood (De La Torre et al. 2021) and sugar pine (Weiss et al. 2022). Current and future studies include comparative population genomic studies to test whether divergent or parallel evolution has occurred in different species; whether natural selection acts primarily on standing genetic variation or on new mutations; and how adaptive evolution has occurred in the different taxa. This work contributes to the study of local adaptation in ecologically important conifer species, and will inform conservation and management practices under climate change.
Active Research Projects:
- Genomic basis of local adaptation among varieties of Ponderosa pine (funding/collaborators: BLM, NAU)

Hybridization and the maintenance of species barriers
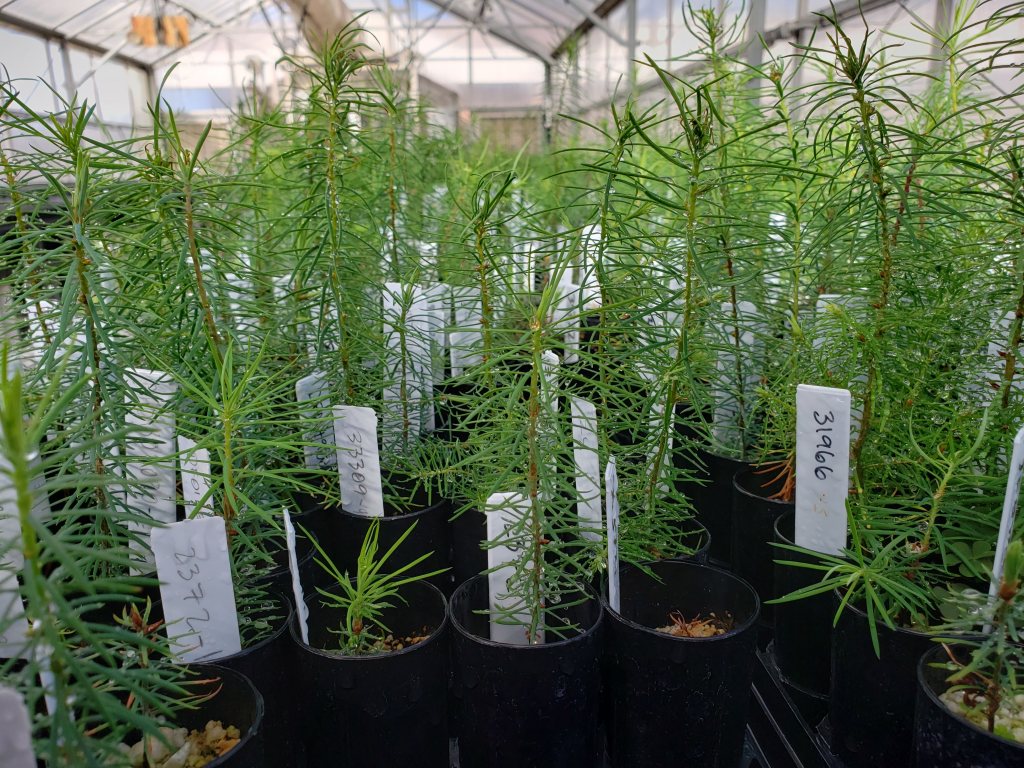

When ecological divergence is driving the evolution of reproductive isolation in natural populations, identifying loci involved in local adaptation is the first step to understand the process of speciation and the maintainance of species identities. In our study in interior spruce, Genome-wide studies, as well as paleoclimatic and ecological niche modeling suggested that despite a long history of hybridization and introgression, dating to at least 21,000 years ago, species identity was maintained by a combination of strong environmental selection acting on a small number of widely distributed genes and reduced current interspecific gene flow (De La Torre et al. 2014 Mol Ecology). Current research in our group aims to understand how different hybridizing tree varieties have evolved different adaptations to contrasting environments (e.g. coastal vs. mountainous).
Active Research Projects:
- Patterns of admixture, introgression, demographic history and phylogenetics in Douglas fir (funding/collaborators: NIFA, NSF, BCMinistry of Forests, UNAM).
Genomics of complex traits
Most traits of interest in plant species have complex inheritance, meaning that a large number of genes of small effect contribute to the phenotype. The understanding of the genomic basis of complex traits is essential to mitigate the devastating effects of disease, drought and cold stress and to develop breeding strategies including genome selection, and marker-assisted breeding. Our recent studies in several conifers such as Loblolly pine, Douglas fir and Sugar pine have identified thousands of gene-trait associations, identifying important genes and pathways in different important traits.
Active Research Projects:
- Genomic, transcriptomic and physiological basis of drought tolerance among varieties of Ponderosa pine (funded by NAU)
- Genomic, transcriptomic, epigenomic and physiological basis of drought and heat tolerance among varieties of Douglas fir (funding/collaborators: NSF, USFS)
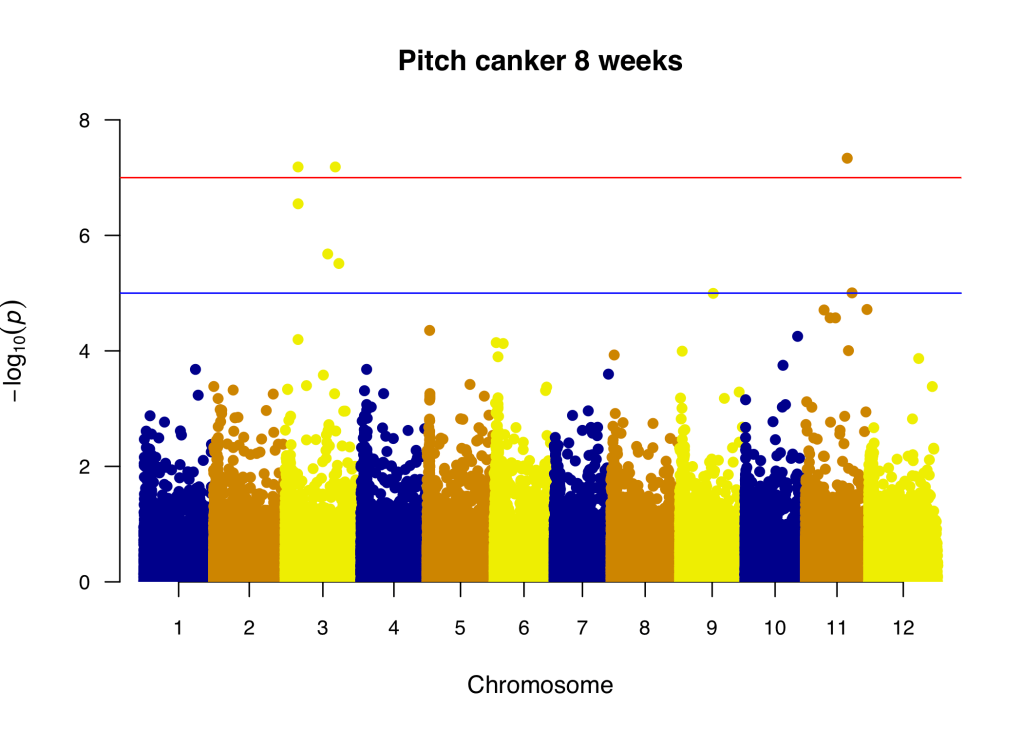
Epigenomics and genomics of disease resistance
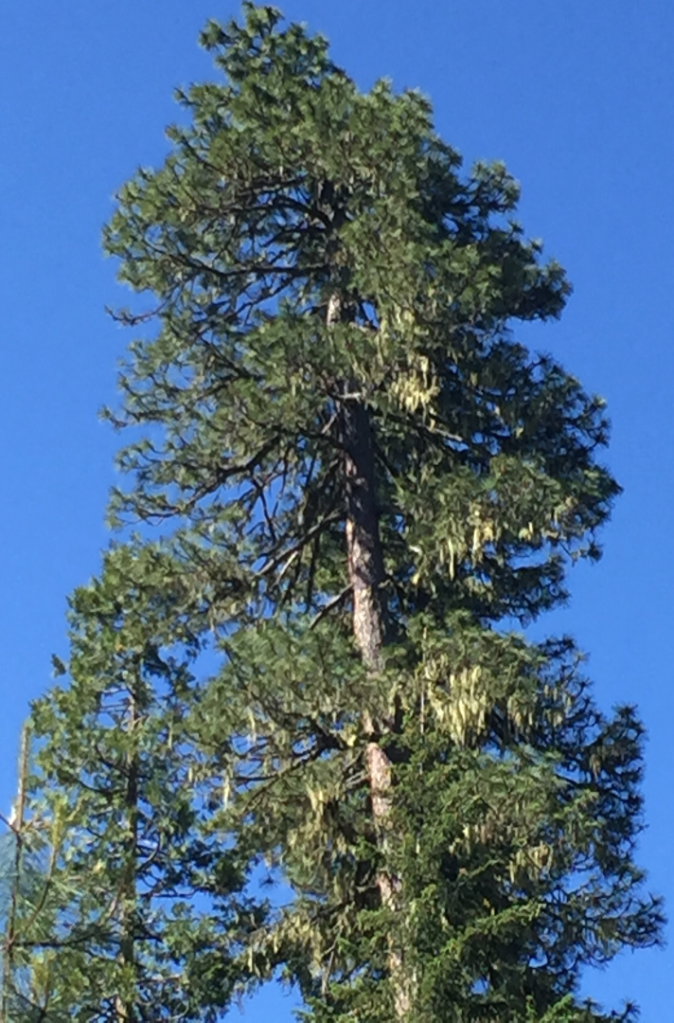

Sugar pine (Pinus lambertiana) growing in Northern California.
Photo taken by Dr. De La Torre in 2016.
Plants possess sophisticated immune responses to defend themselves against pathogens. Recent studies in model plant species suggest DNA methylation may contribute to plant immunity against biotrophic pathogens by regulating the expression of specific defense-related genes. In highly-repetitive, densely-methylated conifer genomes, DNA methylation may play an important role in the response to pathogens, as varying levels of resistance and high levels of phenotypic plasticity and local adaptation are usually observed in natural populations of conifer species, despite the very slow mutation rates (De La Torre et al. 2017). Due to the pervasive expansion of pests and pathogens in the Northern hemisphere as a consequence of climate change, genomic, transcriptomic and epigenomic analyses are key to ensure the trees’ future survival.
Active Research Projects:
- Transcriptomic and Epigenomic responses to white pine blister rust in sugar pine (funding/collaborators: NIFA, NAU SICSS, UConn, USFS)
- Transcriptomic responses to white pine blister rust in whitebark pine (funding/collaborators: NIFA, UConn, USFS)
Comparative Genomics
The recent sequencing of the enormous genomes (20-30 Gbp) of gymnosperms has opened a window to understand the evolution of seed plants. Gymnosperms separated from flowering plants 300 Mya. While flowering plants have gone through successive rounds of species radiations leading to a great diversity of species, gymnosperms have not changed much during evolutionary time (De la Torre et al. 2014 Plant Physiology). In contrast to this apparent “genome stasis”, numerous examples of local adaptation have been reported in gymnosperm species. To answer the question of whether gymnosperms genomes have evolved differently than their sister plant clade of flowering plants, we have studied the patterns of selection and molecular evolution across the genomes of 61 plant species. We found that gymnosperms evolve slower than angiosperms, but present higher substitution rate ratios that may suggest a higher potential for adaptive evolution (De La Torre et al. 2017 Mol Biol Evol). More recently, our group was involved in large collaborative projects to sequence the reference genomes of Populus tremula and P. tremuloides (Li et al. 2018 PNAS), Sequoia sempervirens (Neale et al. 2022), Pinus pinaster (Sterck et al. 2022) and Pinus albicaulis (Neale et al. 2024).
Active Research Projects:
- New assembly, annotation and whole-genome methylation of the 32Gbp Pinus lambertiana genome (funding/collaborators: NIFA, UConn, JHU, USFS)
- de novo transcriptome and genome annotation of the whitebark pine (P. albicaulis) genome (funding/collaborators: NIFA, UConn, JHU, USFS)
- New assembly and annotation of the Douglas fir (P.menziesii) 16Gbp genome (funding/collaborators: USFS, NSF,JHU)
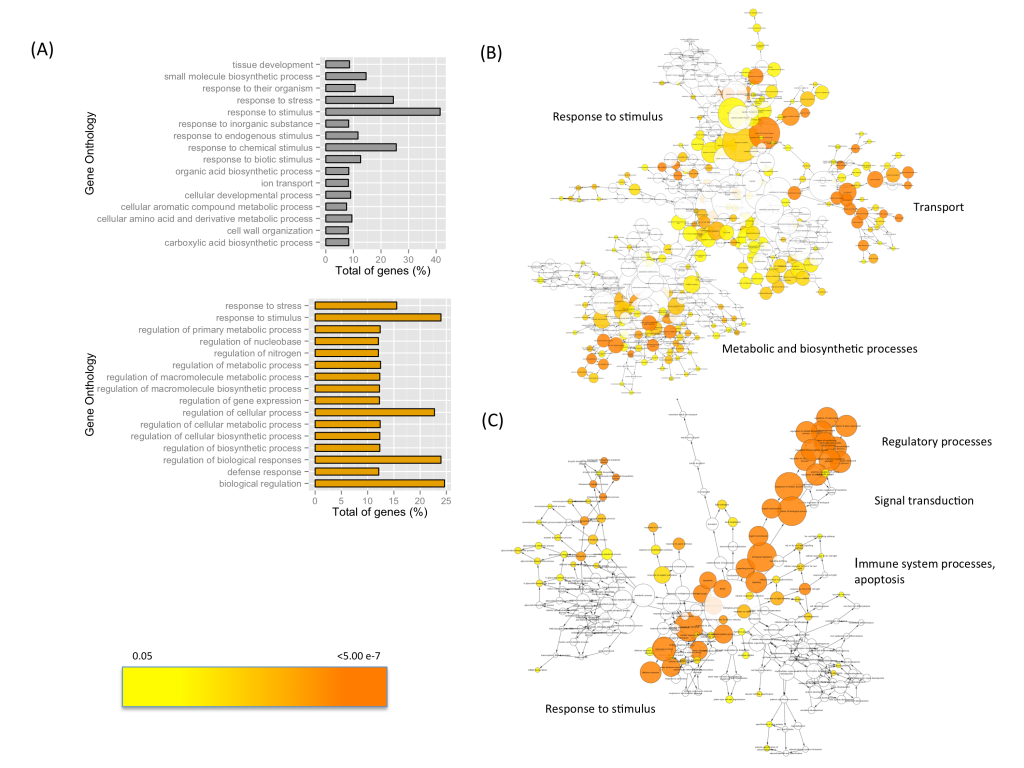
Conservation Genomics
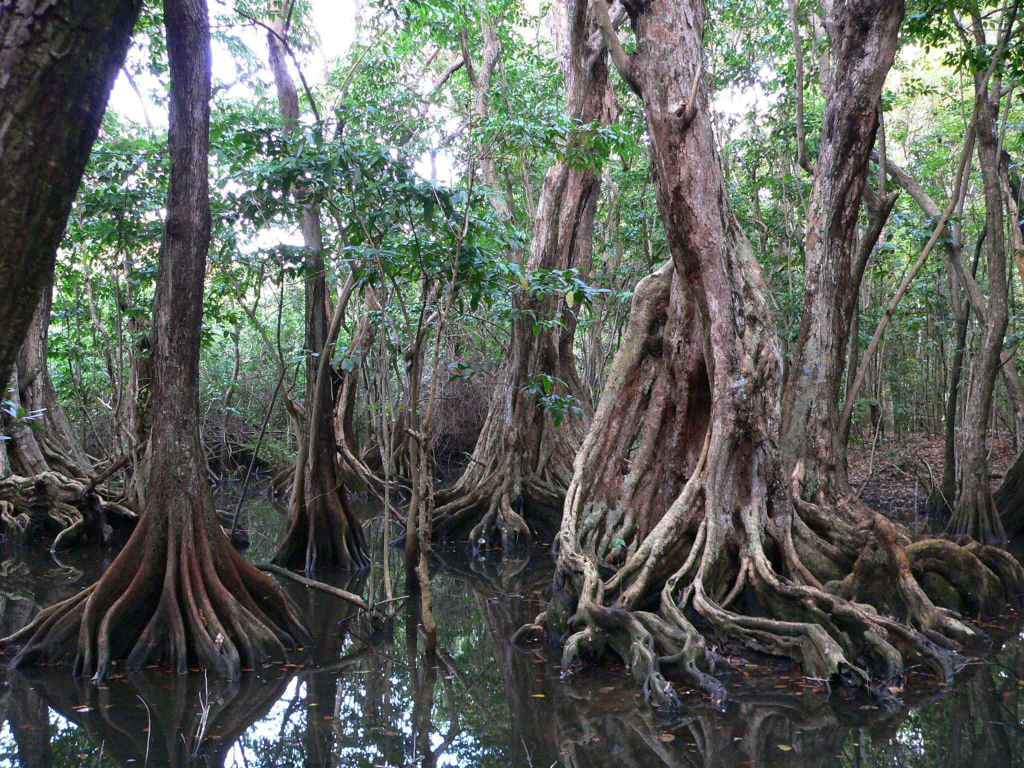

Genomic approaches can be used to help develop long-term conservation and restoration plans for species threatened by fragmentation, overexploitation, invasive species, habitat loss, and climate change. Recent projects in our group, aim to assess the levels of genome-wide diversity in isolated and endangered populations of whitebark pine (Pinus albicaulis) that will inform conservation and restoration of the species. In addition, our group is also assessing the levels of genomic diversity, fine-scale population structure, and salinity resistance in mangrove populations in the Caribbean.
Active Research Projects:
- Population genomics and drought tolerance in Pterocarpus spp. in the Caribbean (funded by USFS).
- Conservation Genomics of whitebark pine (funding/collaborators: NPS, BLM, Parks Canada, USFS)
